Timeline of
Ancient DNA
Ancient DNA research begins with genetic analysis of 140-year-old ancient zebra
1984
1985
Researchers look at 2,400-year-old Egyptian mummies to understand ancient DNA
DNA analysis reveals closest living relatives to now-extinct Tasmanian tiger
1989
1990
The best-selling novel Jurassic Park introduces ancient DNA technology to readers
DNA from extinct giant birds recovered from 3,550 years old museum specimens
1992
1994
DNA of tuberculosis found in 1,000-year-old pre-Columbian Peruvian mummy
First woolly mammoth DNA recovered from Siberian species
1994
1997
90 centuries and 300 generations later, DNA links British caveman to local teacher
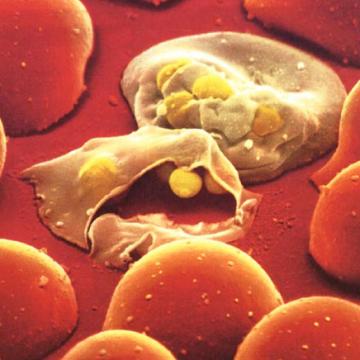
First successful identification of ancient malaria DNA from human remains
1997
1997
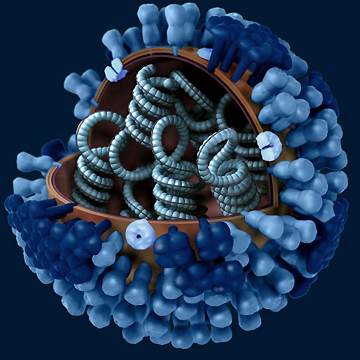
Scientists study ancient virus for answers to the deadly 1918 “Spanish flu” pandemic
DNA from ground sloths’ fossilized dung reveals ecology of lost world
1998
2003
From teosinte to tortillas: Domestication of corn is revealed
Ice cores from Siberia reveal DNA of ancient plants and animals
2003
2005
Scientists recover DNA from 40,000-year-old cave bears and their environment
Paleoindians brought bottle gourds to Americas 10,000 years ago – or did they?
2005
2007
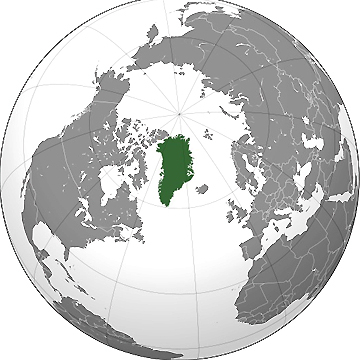
Ice-covered Greenland was a conifer forest half-a-million years ago
Fossilized eggshells yield well-preserved DNA – even in warm climates
2010
2010
Secrets of the Pharaohs: family tree of King Tutankhamen unveiled
Neanderthal genome yields insights into human evolution
2010
2010
Early Greenland settler “Inuk” has whole genome decoded
Cave paintings show true colors of Stone Age horses
2011
2011
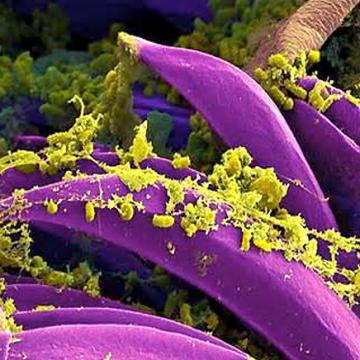
Black Death genome dug out from burials in a London plague cemetery
Fossilized fingertip in Siberia reveals missing human species, the Denisovans
2011
2012
Ancient syphilis genome recovered from newborns deceased in the 16th-17th Century
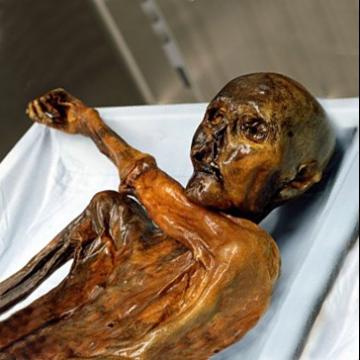
Tyrolean Iceman mummy, “Ötzi,” has whole genome decoded
2012
2013
Medieval leprosy genome compared with modern strains
Mystery of Irish potato famine solved by DNA sequencing 170 years later
2013
2013
Ancient medical kit in shipwreck reveals ingredients of Roman medicine
700,000-year-old horse genome shatters the age record for ancient DNA
2013
2013
DNA in fossilized dental plaque traces human diet from Neolithic through modern times
“Clovis Boy” DNA links ancient boy to American Indians and other Native peoples in the Americas
2014
2014
Genetic studies reveal kiwis are the closest living relatives of the huge elephant birds

Explore Genomic Resources
Our free resource library is packed full of lesson plans, videos, interactive games and other educational content from the National Human Genome Research Institute and our partners.

Read Genomics: Insights
Read articles written by promising researchers about the science they're doing in the lab to inform, educate, and raise awareness about genetics and genomics.