Genomics: Insight

The Laccase Gene Family: Driver of Fungal-Based Bioremediation of Contaminated Soils
Research Question: How does the laccase gene family enable fungi to respond to toxic compounds in contaminated soil and effectively degrade those pollutants?
Significance
Polycyclic aromatic hydrocarbons (PAHs) are an environmental and global health crisis. As of 2025, the Agency for Toxic Substances and Disease Registry recognizes PAHs as primary contaminants in nearly 45% of all active US Superfund sites, where they persist due to their hydrophobic nature (1). The extreme hydrophobicity often makes PAHs difficult for enzymes to target because they don’t dissolve well in water and bind to soil particles. The environmental burden of PAHs directly harms human health. Individuals in highly contaminated zones face a rising global mortality rate from PAH-attributed lung cancer, which has seen an estimated annual increase of 1.14% (2). Furthermore, recent research indicates that women living within four miles of these contaminated sites have a 27% higher risk of being diagnosed with breast cancer compared to those in unpolluted sites (2).
By utilizing fungal species to degrade these toxins, mycoremediation can offer a pathway to reducing these significant health risks. Fungi achieve this by secreting laccases, extracellular enzymes that oxidize toxic substances composed of large amounts of aromatic compounds. These enzymes cause structural instability within the toxin, allowing for biodegradation to occur at a faster rate in contaminated soils. Several fungal species have multiple copies of the laccase gene –all of which perform the same general function while maintaining key differences that increase fungal versatility– including Pleurotus ostreatus, which contains approximately 10-12 different laccase genes (5). This increased copy number results in a larger capacity for Pleurotus ostreatus to produce the enzymes necessary to degrade toxic substances in order to enhance PAH degradation.
Analysis
Figure 1 – Integrated Model of Laccase-Mediated PAH Detoxification and Associated Genomic Regulatory Elements (MRE/XRE)
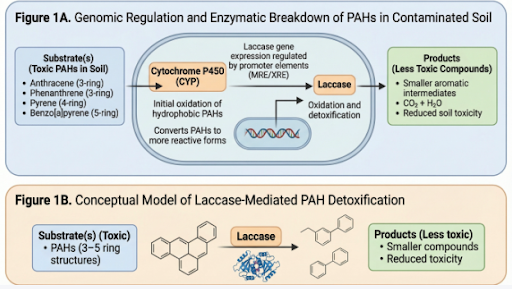
The genomic regions that have the strongest association with PAH degradation in fungi are laccase gene clusters; MRE/XRE (stress-responsive promoters); heterodimeric laccases (such as POXA3); and cytochrome P450 (CYP) genes (11). Laccase gene clusters encode for several different but similar genes which allow the fungus to produce sufficient amounts of an enzyme that is capable of degrading 3-4 ring PAHs (anthracene/pyrene) in contaminated soils at rates of 60-90%. The MRE/XRE promoter regions are areas of DNA that act as switches to increase laccase gene expression. Specifically, PAHs act as a trigger for the XRE, but are influenced distinctly by heavy metals and PAHs. Fungi hve evolved a mechanism where MRE and XRE work together to increase enzyme production when either or both threats are detected. The heterodimeric form of the laccase enzymes increases their stability and efficiency under harsh environmental soil conditions. The CYP genes begin the chemical modification of the large hydrophobic PAHs so that they can be oxidized by laccase. In combination, these genetic systems provide a well-coordinated genomic system that includes regulated laccase gene expression that has the strongest positive association with effective removal of PAHs from contaminated soils.
Figure 2 – Enzyme Comparison for PAH degradation in White-Rot Fungi
Enzyme | Requirements. | Substrate Range | Degradation Examples from Studies | Key Advantages | Key Limitations |
|---|---|---|---|---|---|
Laccase | • Requires only O₂ • Produces H₂O as byproduct • Enhanced by mediators | • 3-5 ring PAHs • Phenolic & non-phenolic compounds • Diverse aromatic substrates | • Anthracene: 22% without mediators, 60% with ABTS (6h) (12) • Enhanced by natural mediators like 4-hydroxybenzoic acid (13) • P. ostreatus: 74.2% benzo[a]pyrene with copper, 78.7% with ABTS (14) | • Broad substrate versatility (12, 13) • Simple oxygen-only requirement(12) • Works on both phenolic and non-phenolic PAHs (13) • Can be enhanced by natural mediators (13) | • Performance depends on mediator presence (12) • Some high MW PAHs require mediators (12) • Variable activity across conditions |
Lignin Peroxide (LiP) | • Requires H₂O₂
| • Primarily non-phenolic PAHs • Aromatic compounds | • Can oxidize benzo[a]pyrene • Effective on recalcitrant compounds | • Can oxidize non-phenolic substrates • High oxidation potential • Attacks aromatics via free radicals | • Requires H₂O₂ (15) • Less commonly produced than MnP • Sensitive to environmental conditions |
Manganese Peroxide (MnP) | • Requires H₂O₂ + Mn²⁺ • Mn³⁺ acts as a mediator
| • Phenolic PAHs • Various recalcitrant compounds | • Most common ligninolytic enzyme in white-rot fungi • 100 mg/L benzo[a]pyrene converted in 24h with 1 U/mL MnP • Present in almost all wood-colonizing white-rot fungi (16) | • Most abundant enzyme in white-rot fungi (16) • Mn³⁺ diffuses to reach embedded pollutants • Produced by almost all white-rot species (16) | • Requires both H₂O₂ and Mn²⁺ • Primarily oxidizes phenolic compounds • Indirect oxidation mechanism |
Versatile Peroxidase (VP) | • Works with/without Mn²⁺ • Requires H₂O₂ | • 3-5 ring PAHs • Both phenolic and non-phenolic • Polymeric substrates | • Found in P. ostreatus as part of the 9-gene MnP/VP family (17, 18) • Directly oxidizes polymeric substrates without mediators (3) • Mnp4 is predominantly expressed VP in Mn²⁺-deficient media (18) | • Hybrid of LiP and MnP functions (3, 17) • Present in P. ostreatus (17, 18) • Can oxidize substrates with/without Mn²⁺ (3, 17) • Directly oxidizes polymers (3) | • Part of a complex gene family with redundancy (3, 17) • Still requires H₂O₂ (3) • Differentially regulated by Mn²⁺ (16, 18) |
Among the oxidative enzymes secreted by the white-rot fungi, laccase distinguishes itself from the rest as the most promising bioremediation tool (Table 2). This is due to laccase’s broad substrate versatility and simple catalytic requirements, requiring only molecular oxygen while producing water as a byproduct, in contrast to peroxidases (MnP, LiP, VP), which require hydrogen peroxide (12, 13). Studies demonstrate laccase achieved 60% anthracene degradation with ABTS mediator in 6 hours, while Pleurotus ostreatus degraded 78.7% of benzo[a]pyrene with ABTS (12, 14). P. ostreatus has a unique ligninolytic system with nine MnP/VP-encoding genes, including versatile peroxidase, which combines LiP and MnP functions to oxidize both phenolic and non-phenolic substrates (3, 17,18). Laccase's combination of substrate versatility, oxygen-only requirements, and enhancement through natural mediators positions it as highly adaptable for mycoremediation applications (12,13).
Functional Significance:
The laccase gene family is one of the most effective in breaking down toxic PAHs in the soil. They have a broad substrate range, which means they can act on a large variety of PAHs –PAHs are classified into 3-4 ring and 5 ring– and have minimal requirements to work. Species like Trametes versicolor, Cerrena unicolor, and Phlebia serialis are the main degraders that can break down 3 to 4-ring PAHs (4). Fungal species utilize different regulatory mechanisms to control laccase production depending on environmental triggers in the surrounding soil. Examples of species that possess specialized promoter regions (MRE/XRE) are Pleurotus Ostreatus and Agaricus bisporus, which trigger laccase expression to heavy metals or PAHs. These regulated laccases are capable of degrading smaller 3-4 ring PAHs at high efficiencies, while Cytochrome P450 is needed to degrade 5-ring PAHs because they are larger and more hydrophobic (4). The main hurdle with breaking down 5-ring PAHs is that they are extremely hydrophobic. To overcome this barrier, the Cytochrome P450 monooxygenases activate the hydrophobic PAHs by hydroxylating and polarizing the molecules, which makes them ready for enzymatic oxidation. A large-scale study using Pleurotus Ostreatus in a pilot biopile (a controlled pile of contaminated soil treated to enhance breakdown of pollutants) found that this combination, where Cytochrome P450 primes the molecule for laccase oxidation, was able to reduce the toxic soil PAH levels by ~98%, finding a reduction in toxic PAHs from 6,469 to 112 mg/kg (4). This study shows that laccase-focused mycoremediation is effective, scalable, and highly effective. Other enzymes like manganese peroxidases also have roles in breaking down particular PAHs or in extreme soil conditions, but laccase is still the most effective due to its minimal requirements and broad substrate range (9).
The laccase gene family is one of the most effective in breaking down toxic PAHs in the soil. They have a broad substrate range, which means they can act on a large variety of PAHs and have minimal requirements to work.
Conclusion
The laccase gene family functions as an integrated genomic network that bridges the gap between environmental sensors and toxin degradation. Through the use of promoter regions (MRE/XRE) that are triggered by environmental factors, fungi like the P. ostreatus can detect PAHs, triggering degradation of 3-4 ring PAHs. With the addition of CYPs, which act as a primer for 5-ring PAHs, breaking down their extreme hydrophobicity, so that the laccase can complete its oxidative process. This coordinated response shows a ~98% reduction in toxic soil PAHs. Ultimately, laccase-focused mycoremediation can be a scalable, highly effective solution that succeeds through a coordinated genetic adaptive response against harmful environmental pollutants. As scientific discoveries continue, the potential applications of these mushrooms could offer a scalable solution for mitigating the global impact of industrial PAH contamination, ultimately protecting public health and restoring ecosystems.
References
- Polycyclic Aromatic Hydrocarbons (PAHs) | Public Health Statement | ATSDR. (2026). Cdc.gov. https://wwwn.cdc.gov/TSP/PHS/PHS.aspx?phsid=120&toxid=25
- Roldan, B. (2026). New Research Links Aggressive Breast Cancers to Superfund Sites. Breastcancer.org. https://www.breastcancer.org/news/superfund-sites-aggressive-cancers
- KAMITSUJI, H., WATANABE, T., HONDA, Y., & KUWAHARA, M. (2005). Direct oxidation of polymeric substrates by multifunctional manganese peroxidase isoenzymes from Pleurotus ostreatus without redox mediators. Biochemical Journal, 386(2), 387–393. https://doi.org/10.1042/bj20040968
- Di Gregorio, S., Becarelli, S., Siracusa, G., Ruffini Castiglione, M., Petroni, G., Masini, G., Gentini, A., de Lima e Silva, M. R., & Lorenzi, R. (2016). Pleurotus ostreatus spent mushroom substrate for the degradation of polycyclic aromatic hydrocarbons: the case study of a pilot dynamic biopile for the decontamination of a historically contaminated soil. Journal of Chemical Technology & Biotechnology, 91(6), 1654–1664. https://doi.org/10.1002/jctb.4936
- Jiao, X., Li, G., Wang, Y., Nie, F., Cheng, X., Abdullah, M., Lin, Y., & Cai, Y. (2018). Systematic Analysis of the Pleurotus ostreatus Laccase Gene (PoLac) Family and Functional Characterization of PoLac2 Involved in the Degradation of Cotton-Straw Lignin. Molecules, 23(4), 880. https://doi.org/10.3390/molecules23040880
- Aza, P., & Camarero, S. (2023). Fungal Laccases: Fundamentals, Engineering and Classification Update. Biomolecules, 13(12), 1716. https://doi.org/10.3390/biom13121716
- Paweena Thongkred, Pongtharin Lotrakul, Sehanat Prasongsuk, Imai, T., & Hunsa Punnapayak. (2011). ScienceAsia, 37(3), 225–225. https://doi.org/10.2306/scienceasia1513-1874.2011.37.225
- Yu, J., Lai, J., Neal, B. M., White, B. J., Banik, M. T., & Dai, S. Y. (2023). Genomic Diversity and Phenotypic Variation in Fungal Decomposers Involved in Bioremediation of Persistent Organic Pollutants. Journal of Fungi, 9(4), 418. https://doi.org/10.3390/jof9040418
- Dong, C.-D., Tiwari, A., Anisha, G. S., Chen, C.-W., Singh, A., Haldar, D., Patel, A. K., & Singhania, R. R. (2023). Laccase: A potential biocatalyst for pollutant degradation. Environmental Pollution, 319, 120999. https://doi.org/10.1016/j.envpol.2023.120999
- Park, H., Min, B., Jang, Y., Kim, J., Lipzen, A., Sharma, A., Andreopoulos, B., Johnson, J., Riley, R., Spatafora, J. W., Henrissat, B., Kim, K. H., Grigoriev, I. V., Kim, J.-J., & Choi, I.-G. (2019). Comprehensive genomic and transcriptomic analysis of polycyclic aromatic hydrocarbon degradation by a mycoremediation fungus, Dentipellis sp. KUC8613. Applied Microbiology and Biotechnology, 103(19), 8145–8155. https://doi.org/10.1007/s00253-019-10089-6
- Bezalel, L., Hadar, Y., & Cerniglia, C. E. (1997). Enzymatic Mechanisms Involved in Phenanthrene Degradation by the White Rot Fungus Pleurotus ostreatus. Applied and Environmental Microbiology, 63(7), 2495–2501. https://doi.org/10.1128/aem.63.7.2495-2501.1997
- Pickard, M. A., Roman, R., Tinoco, R., & Vazquez-Duhalt, R. (1999). Polycyclic Aromatic Hydrocarbon Metabolism by White Rot Fungi and Oxidation by Coriolopsis gallica UAMH 8260 Laccase. Applied and Environmental Microbiology, 65(9), 3805–3809. https://doi.org/10.1128/aem.65.9.3805-3809.1999
- Johannes, C., & Majcherczyk, A. (2000). Natural Mediators in the Oxidation of Polycyclic Aromatic Hydrocarbons by Laccase Mediator Systems. Applied and Environmental Microbiology, 66(2), 524–528. https://doi.org/10.1128/aem.66.2.524-528.2000
- Bhattacharya, S., Das, A., Prashanthi, K., Palaniswamy, M., & Angayarkanni, J. (2014). Mycoremediation of Benzo[a]pyrene by Pleurotus ostreatus in the presence of heavy metals and mediators. 3 Biotech, 4(2), 205–211. https://doi.org/10.1007/s13205-013-0148-y
- Chatterjee, P., Kouzi, S. A., Pezzuto, J. M., & Hamann, M. T. (2000). Biotransformation of the Antimelanoma Agent Betulinic Acid by Bacillus megaterium ATCC 13368. Applied and Environmental Microbiology, 66(9), 3850–3855. https://doi.org/10.1128/aem.66.9.3850-3855.2000
- Cohen, R., Yarden, O., & Hadar, Y. (2002). Lignocellulose Affects Mn 2+ Regulation of Peroxidase Transcript Levels in Solid-State Cultures of Pleurotus ostreatus. Applied and Environmental Microbiology, 68(6), 3156–3158. https://doi.org/10.1128/aem.68.6.3156-3158.2002
- Sekan, A. S., Myronycheva, O. S., Karlsson, O., Gryganskyi, A. P., & Blume, Y. B. (2019). Green potential of Pleurotus spp. in biotechnology. PeerJ, 7, e6664. https://doi.org/10.7717/peerj.6664
- Joyner, J., Wanless, D., Sinigalliano, C. D., & Lipp, E. K. (2014). Use of Quantitative Real-Time PCR for Direct Detection of Serratia marcescens in Marine and Other Aquatic Environments. Applied and Environmental Microbiology, 80(5), 1679–1683. https://doi.org/10.1128/aem.02755-13
About the Author