A Brief History
From Mendel to the Human Genome Project
Gregor Mendel, the father of modern genetics, presents his research on experiments in plant hybridization
1865
1869
Friedrich Miescher identifies "nuclein," DNA with associated proteins, from cell nuclei
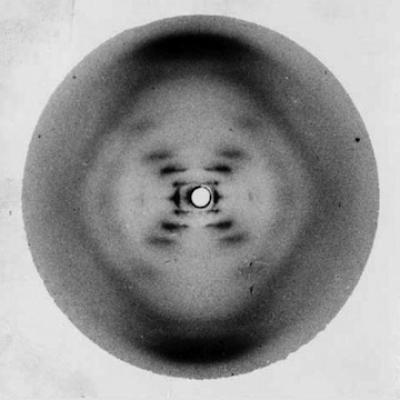
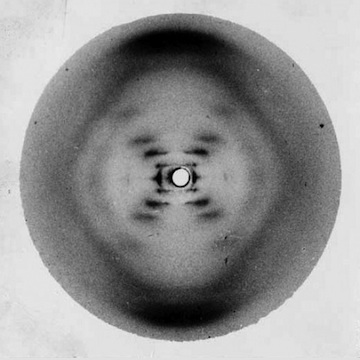
Rosalind Franklin creates Photograph 51, showing a distinctive pattern that indicates the helical shape of DNA
1952
1953
James Watson and Francis Crick discover the double helix structure of DNA
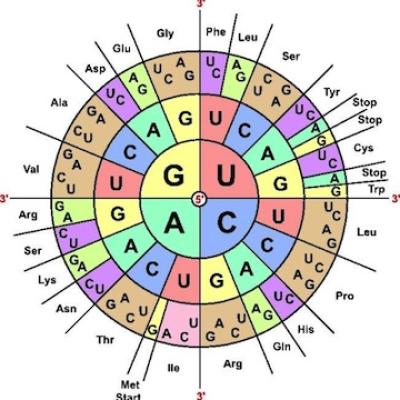
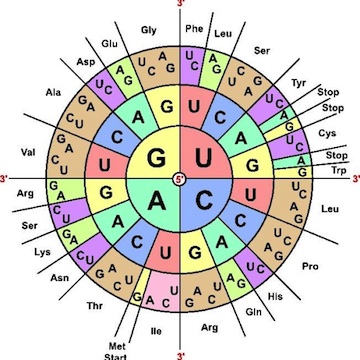
Marshall Nirenberg cracks the genetic code for protein synthesis
1961
1977
Frederick Sanger develops rapid DNA sequencing technique
First genetic disease mapped, Huntington’s Disease
1983
1983
Invention of polymerase chain reaction (PCR) technology for amplifying DNA
Cystic Fibrosis gene mutation identified
1989
1990
First evidence provided for the existence of the BRCA1 gene
The Human Genome Project begins
1990
1995
Haemophilus influenzae becomes first bacterium genome sequenced
“Bermuda Principles” drafted for Human Genome Project free data access
1996
1998
Celera Genomics Corporation founded for sequencing the human genome
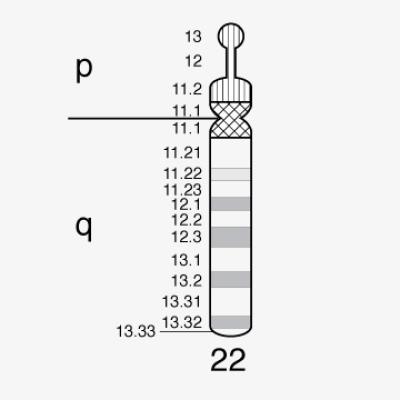
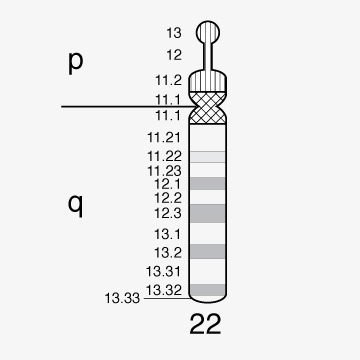
Chromosome 22 first human chromosome to be decoded
1999
2000
Genome sequence of model organism fruit fly reported
First draft of the human genome released
2001
2002
Mouse becomes first mammalian research organism with decoded genome
Human Genome Project completion announced
2003

Explore Genomic Resources
Our free resource library is packed full of lesson plans, videos, interactive games and other educational content from the National Human Genome Research Institute and our partners.

Read Genomics: Insights
Read articles written by promising researchers about the science they're doing in the lab to inform, educate, and raise awareness about genetics and genomics.